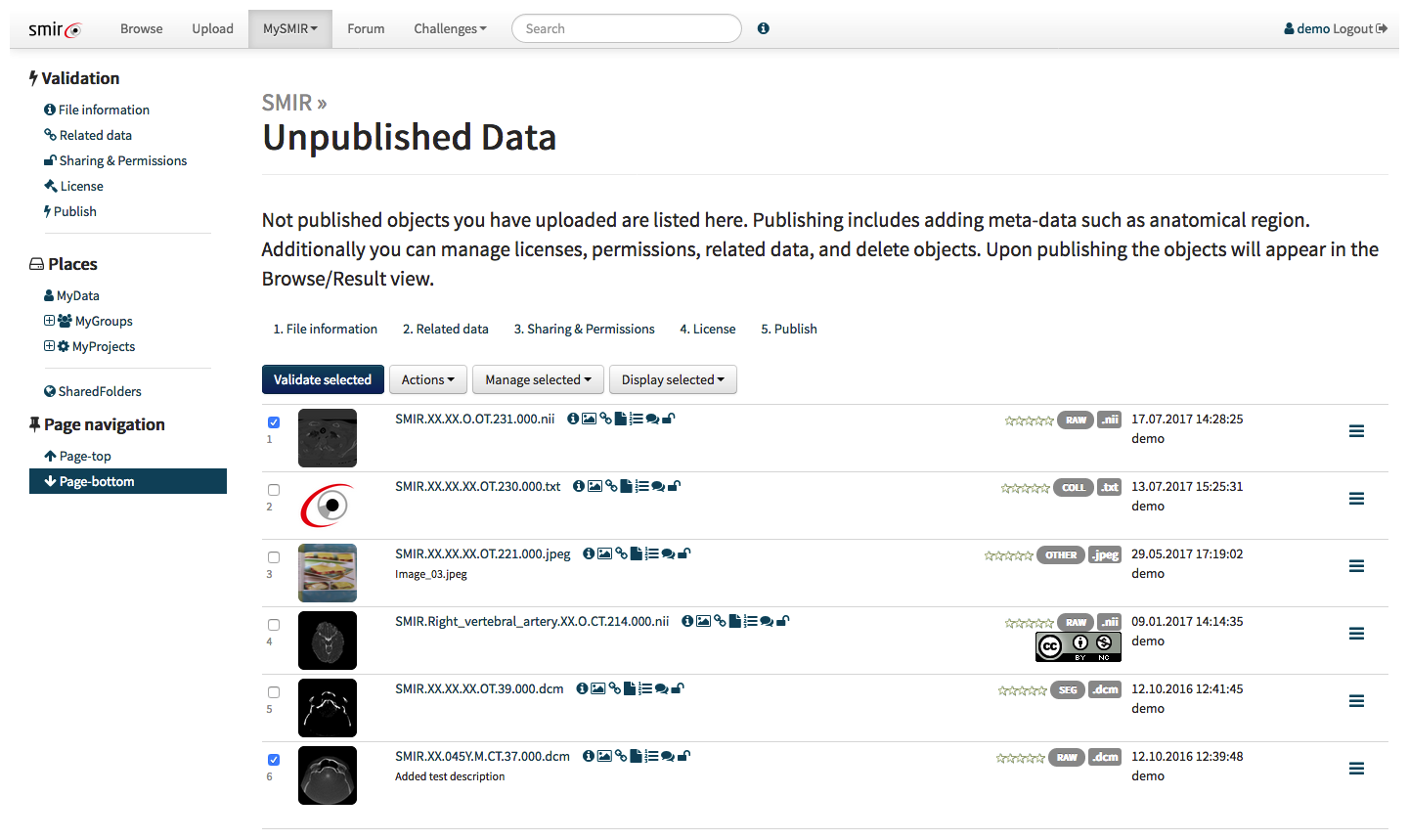
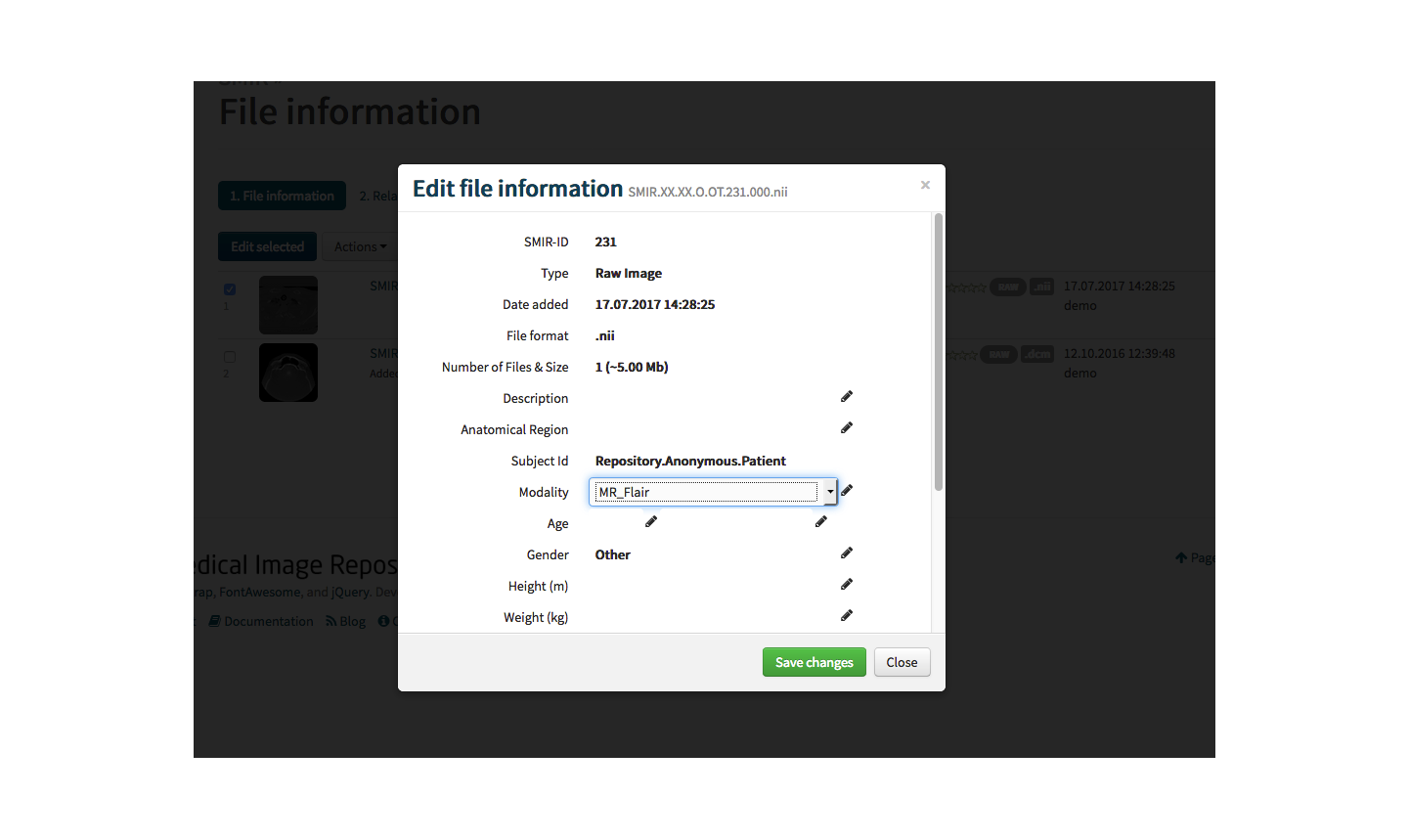
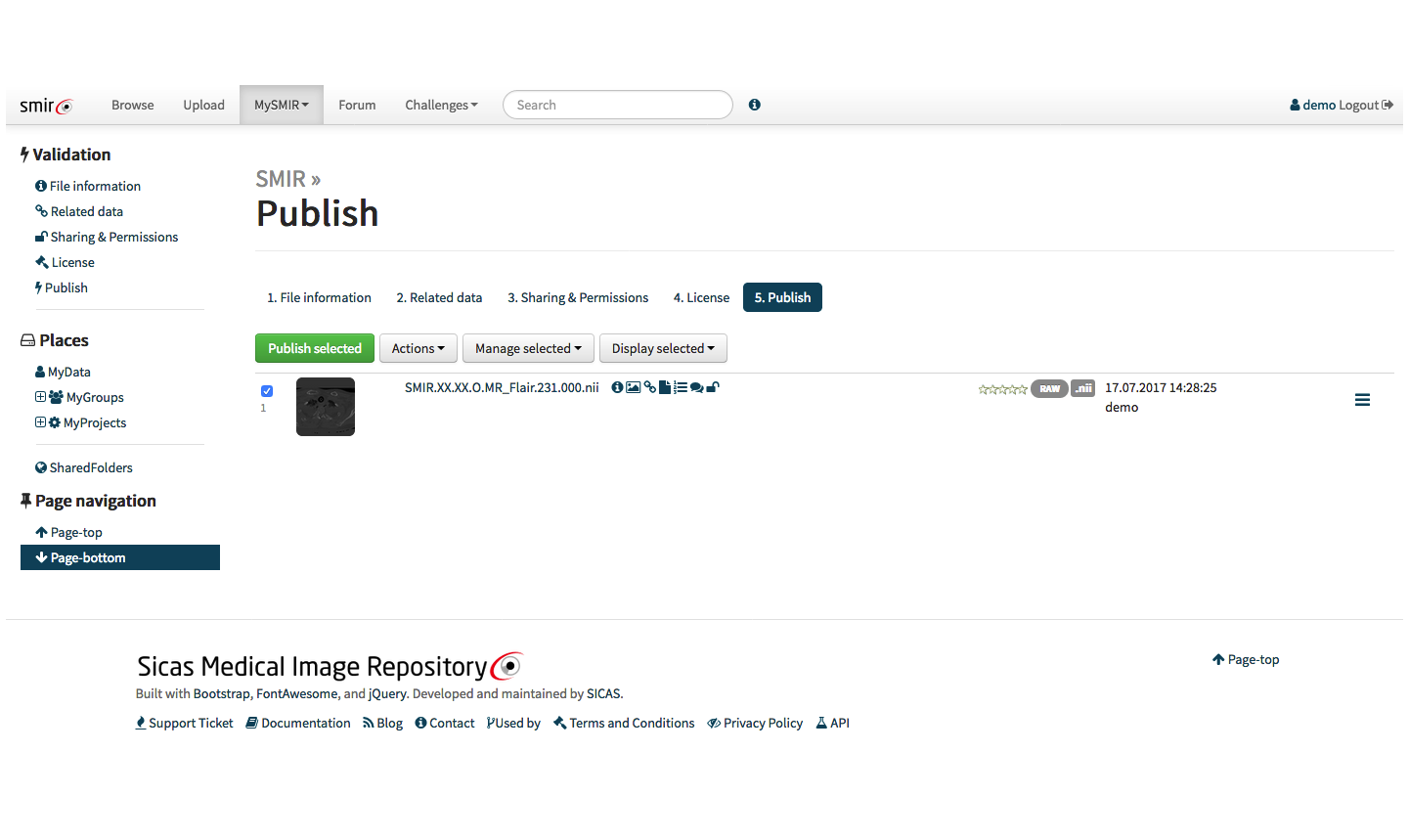
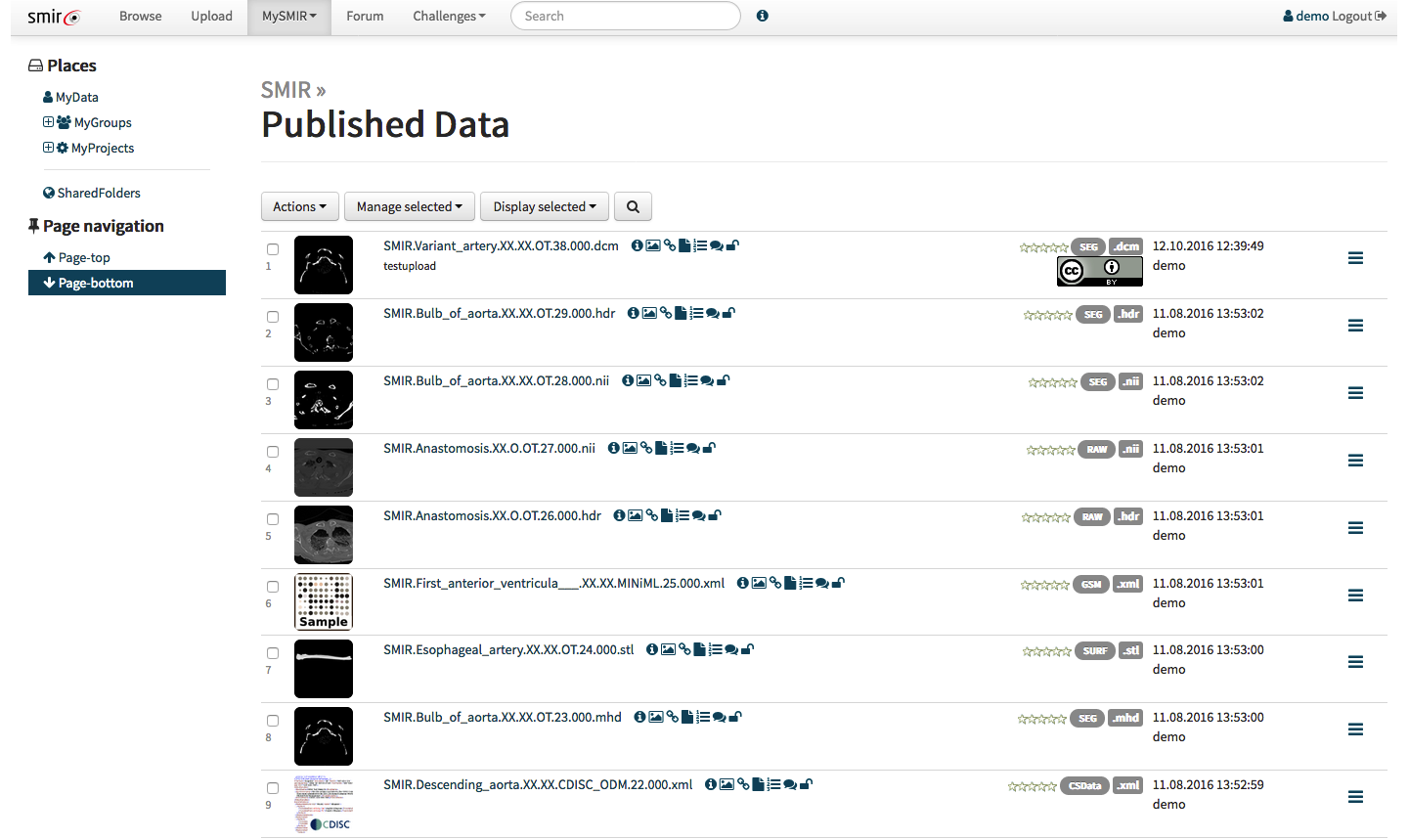
With the API
Please consider developing on the demo (https://demo.smir.ch) server before you interact with the live system.
API Reference
Python 3 API Connector
Scala (JAVA) API Connector
Chunked file upload
Python code
## import libs
from pathlib import Path
from vsdConnect import connect
## connect to demo
api = connect.VSDConnecter()
## define filepath
fp = Path('C:' + os.sep, 'test', 'test.nii')
## Define chunk size to eg. 8 MB if you dont want to use the default 4MB
chunk = 1024 * 4096 * 2
## upload using the chunkFileUpload
obj = api.chunkFileUpload(fp, chunksize = chunk)
## print the selfUrl of the generated object
print(obj.selfUrl)
Console output
uploading part 1 of 3
uploaded part 1 of 3
uploading part 2 of 3
uploaded part 2 of 3
uploading part 3 of 3
uploaded part 3 of 3
https://demo.smir.ch/api/objects/1
Download a folder
Python code
from pathlib import Path
from vsdConnect import connect
# connect to the demo API
api = connectVSD.VSDConnecter()
## search for the test folder and retrieve the folder object
folder = api.getFolderByName('One')
for obje in folder.containedObjects:
## get each object in the folder
obj = api.getObject(obje.selfUrl)
## Download directory
home_dir = Path('/Home/User/Downloads')
obj.download(api, working_dir=home_dir,)